LaminDB allows you to query, trace, and validate datasets and models at scale. You get context & memory through a lineage-native lakehouse that supports bio-formats, registries & ontologies while feeling as simple as a file system.
Agent? llms.txt
Why?
(1) Reproducing, tracing & understanding how datasets, models & results are created is critical to quality R&D. Without context, humans & agents make mistakes and cannot close feedback loops across data generation & analysis. Without memory, compute & intelligence are wasted on fragmented, non-compounding tasks — LLM context windows are small.
(2) Training & fine-tuning models with thousands of datasets — across LIMS, ELNs, orthogonal assays — is now a primary path to scaling R&D. But without queryable & validated data or with data locked in organizational & infrastructure silos, it leads to garbage in, garbage out or is quite simply impossible.
Imagine building software without git or pull requests: an agent's actions would be impossible to verify. While code has git and tables have dbt/warehouses, biological data has lacked a framework for managing its unique complexity.
LaminDB fills the gap.
It is a lineage-native lakehouse that understands bio-registries and formats (AnnData, .zarr, …) based on the established open data stack:
Postgres/SQLite for metadata and cross-platform storage for datasets.
By offering queries, tracing & validation in a single API, LaminDB provides the context & memory to turn messy, agentic biological R&D into a scalable process.
How?
- lineage → track inputs & outputs of notebooks, scripts, functions & pipelines with a single line of code
- lakehouse → manage, monitor & validate schemas for standard and bio formats; query across many datasets
- FAIR datasets → validate & annotate
DataFrame,AnnData,SpatialData,parquet,zarr, … - LIMS & ELN → programmatic experimental design with bio-registries, ontologies & markdown notes
- unified access → storage locations (local, S3, GCP, …), SQL databases (Postgres, SQLite) & ontologies
- reproducible → auto-track source code & compute environments with data & code versioning
- change management → branching & merging similar to git, plan management for agents
- zero lock-in → runs anywhere on open standards (Postgres, SQLite,
parquet,zarr, etc.) - scalable → you hit storage & database directly through your
pydataor R stack, no REST API involved - simple → just
pip installfrom PyPI orinstall.packages('laminr')from CRAN - distributed → zero-copy & lineage-aware data sharing across infrastructure (databases & storage locations)
- integrations → git, nextflow, vitessce, redun, and more
- extensible → create custom plug-ins based on the Django ORM, the basis for LaminDB's registries
GUI, permissions, audit logs? LaminHub is a collaboration hub built on LaminDB similar to how GitHub is built on git.
Who?
Scientists and engineers at leading research institutions and biotech companies, including:
- Industry → Pfizer, Altos Labs, Ensocell Therapeutics, ...
- Academia & Research → scverse, DZNE (National Research Center for Neuro-Degenerative Diseases), Helmholtz Munich (National Research Center for Environmental Health), ...
- Research Hospitals → Global Immunological Swarm Learning Network: Harvard, MIT, Stanford, ETH Zürich, Charité, U Bonn, Mount Sinai, ...
From personal research projects to pharma-scale deployments managing petabytes of data across:
| entities | OOMs |
|---|---|
| observations & datasets | 10¹² & 10⁶ |
| runs & transforms | 10⁹ & 10⁵ |
| proteins & genes | 10⁹ & 10⁶ |
| biosamples & species | 10⁵ & 10² |
| ... | ... |
To install the Python package with recommended dependencies, use:
pip install lamindbInstall with minimal dependencies.
The lamindb package adds data-science related dependencies, those that come with the [full] extra, see here.
If you want a maximally lightweight install of the lamindb namespace, use:
pip install lamindb-coreThis suffices to support the basic functionality but you will get an ImportError if you're e.g. trying to validate a DataFrame because that requires pandera.
You can browse public databases at lamin.ai/explore. To query laminlabs/cellxgene, run:
import lamindb as ln
db = ln.DB("laminlabs/cellxgene") # a database object for queries
df = db.Artifact.to_dataframe() # a dataframe listing datasets & modelsTo get a specific dataset, run:
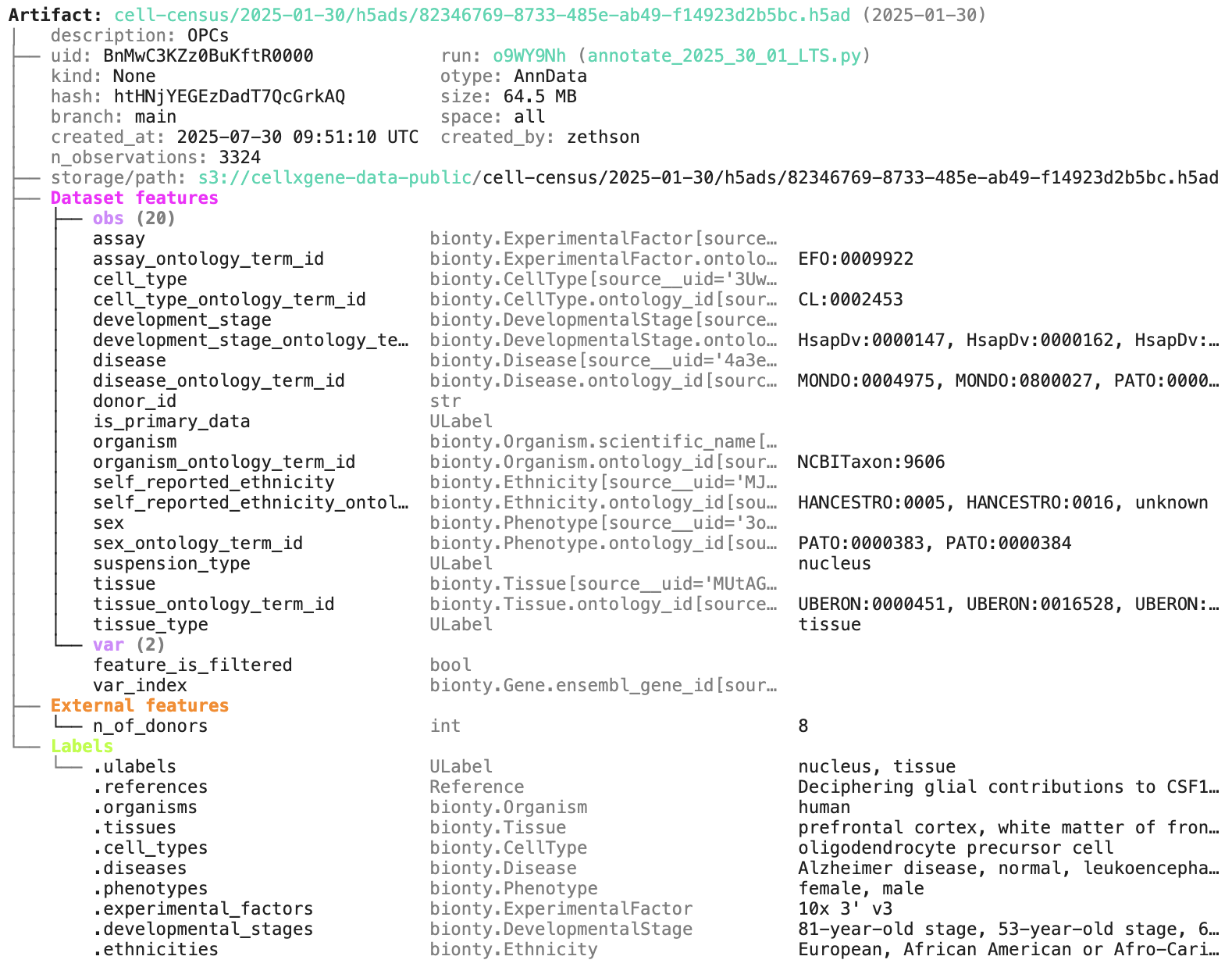
artifact = db.Artifact.get("BnMwC3KZz0BuKftR") # a metadata object for a dataset
artifact.describe() # describe the context of the datasetAccess the content of the dataset via:
local_path = artifact.cache() # return a local path from a cache
adata = artifact.load() # load object into memory
accessor = artifact.open() # return a streaming accessorYou can query by biological entities like Disease through plug-in bionty:
alzheimers = db.bionty.Disease.get(name="Alzheimer disease")
df = db.Artifact.filter(diseases=alzheimers).to_dataframe()You can create a LaminDB instance at lamin.ai and invite collaborators. To connect to an existing instance, run:
# log into LaminHub
lamin login
# then either
lamin connect account/name # connect globally in your environment
# or
lamin connect --here account/name # connect in your current development directoryIf you prefer to init a new instance instead (no login required), run:
lamin init --storage ./quickstart-data --modules biontyFor more configuration, read: docs.lamin.ai/setup.
On the terminal and in a Python session, LaminDB will now auto-connect.
To save a file or folder via the API:
import lamindb as ln
# → connected lamindb: account/instance
open("sample.fasta", "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact("sample.fasta", key="sample.fasta").save() # save datasetTo save a file or folder via the CLI, run:
lamin save sample.fasta --key sample.fastaTo load an artifact via the CLI into a local cache, run:
lamin load --key sample.fastaRead more about the CLI: docs.lamin.ai/cli.
To create a dataset while tracking source code, inputs, outputs, logs, and environment:
import lamindb as ln
# → connected lamindb: account/instance
ln.track() # track code execution
open("sample.fasta", "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact("sample.fasta", key="sample.fasta").save() # save dataset
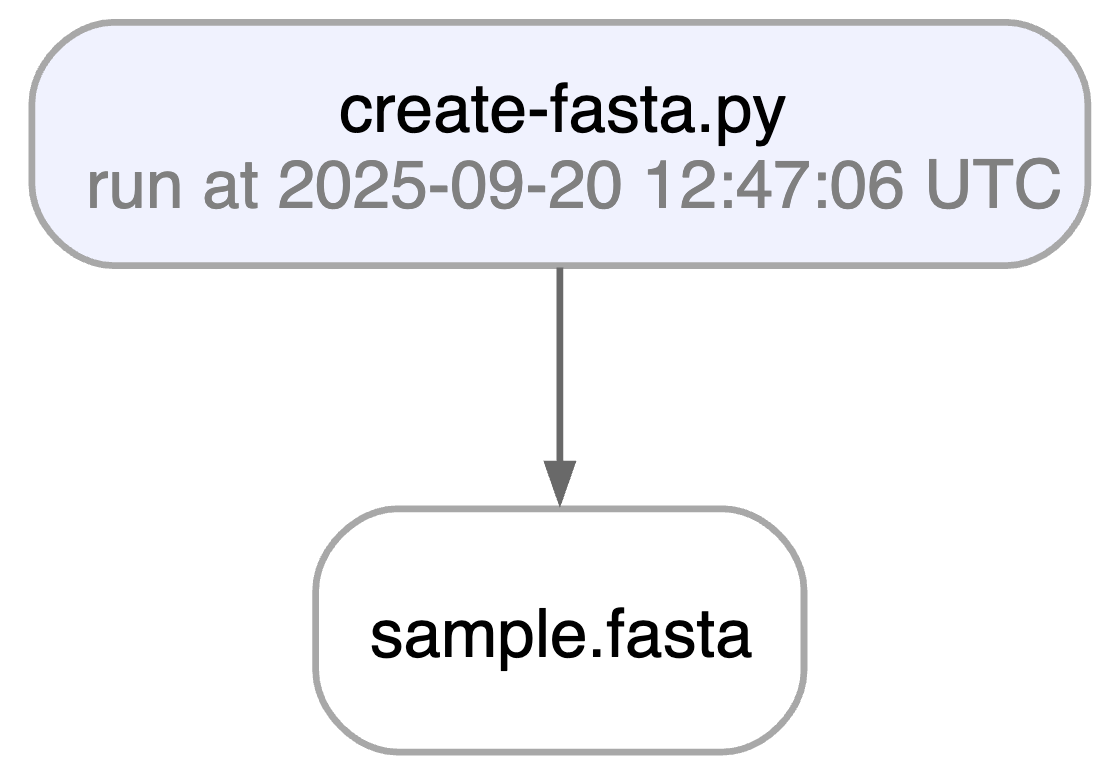
ln.finish() # mark run as finishedRunning this snippet as a script (python create-fasta.py) produces the following data lineage:
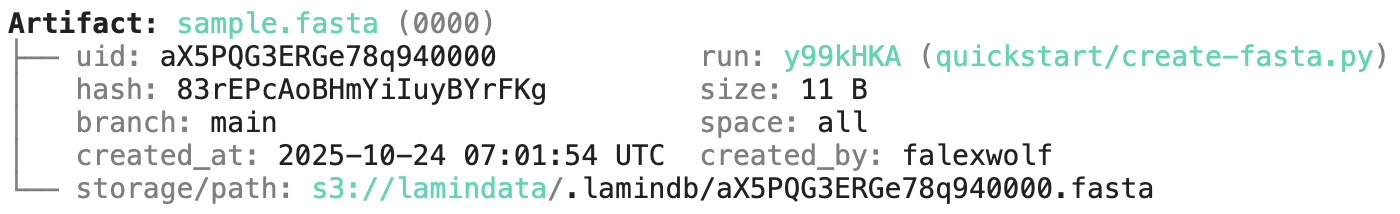
artifact = ln.Artifact.get(key="sample.fasta") # get artifact by key
artifact.describe() # context of the artifact
artifact.view_lineage() # fine-grained lineageAccess run & transform.
run = artifact.run # get the run object
transform = artifact.transform # get the transform object
run.describe() # context of the run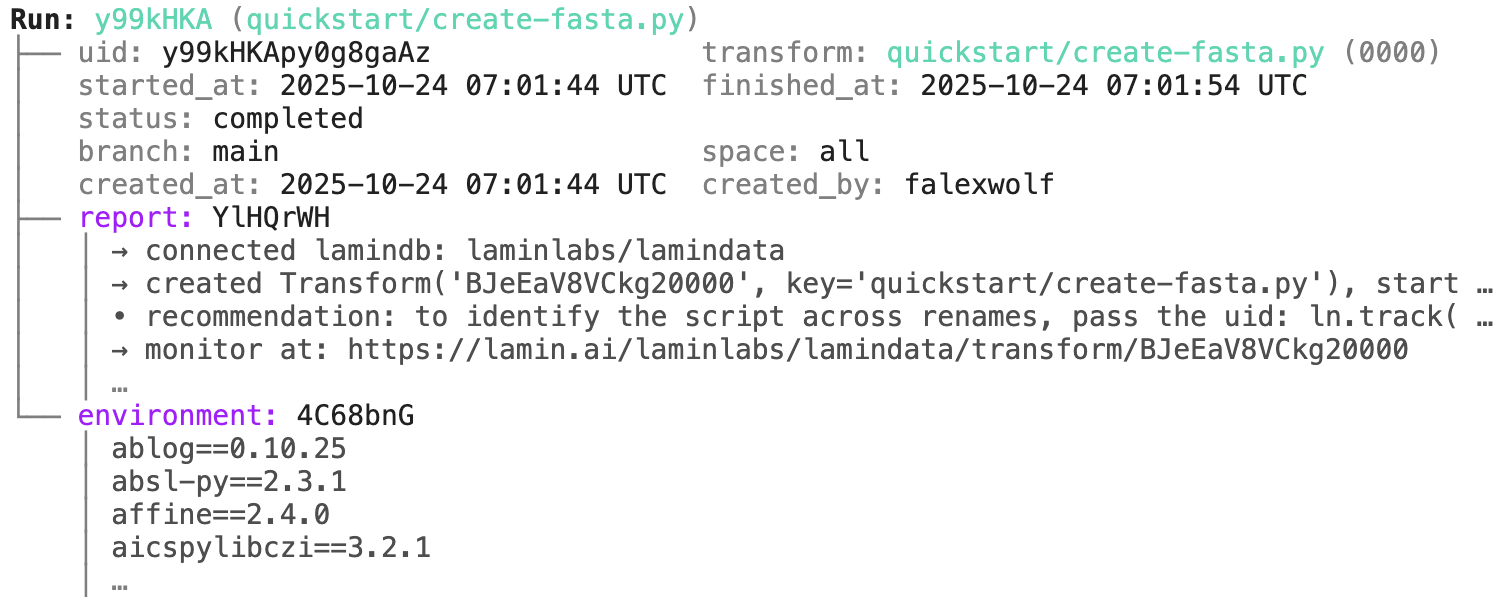
transform.describe() # context of the transform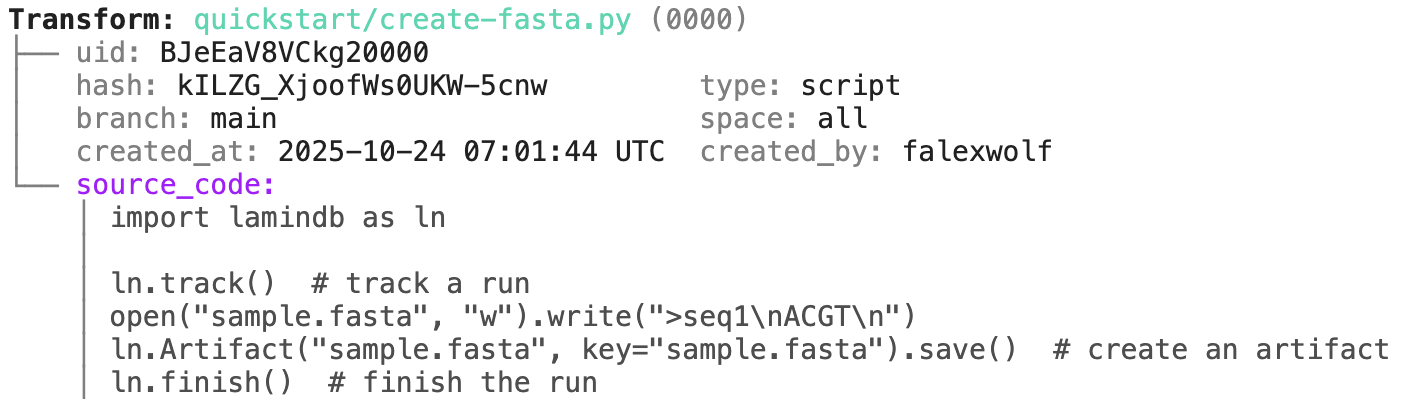
15 sec video.
Track a project or an agent plan.
Pass a project/artifact to ln.track(), for example:
ln.track(project="My project", plan="./plans/curate-dataset-x.md")Note that you have to create a project or save the agent plan in case they don't yet exist:
# create a project with the CLI
lamin create project "My project"
# save an agent plan with the CLI
lamin save /path/to/.cursor/plans/curate-dataset-x.plan.md
lamin save /path/to/.claude/plans/curate-dataset-x.mdOr in Python:
ln.Project(name="My project").save() # create a project in PythonYou can achieve the same traceability for functions & workflows:
import lamindb as ln
@ln.flow()
def create_fasta(fasta_file: str = "sample.fasta"):
open(fasta_file, "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact(fasta_file, key=fasta_file).save() # save dataset
if __name__ == "__main__":
create_fasta()Beyond what you get for scripts & notebooks, this automatically tracks function & CLI params and integrates well with established Python workflow managers: docs.lamin.ai/track. To integrate advanced bioinformatics pipeline managers like Nextflow, see docs.lamin.ai/pipelines.
A richer example.
Here is an automatically generated re-construction of the project of Schmidt et al. (Science, 2022):
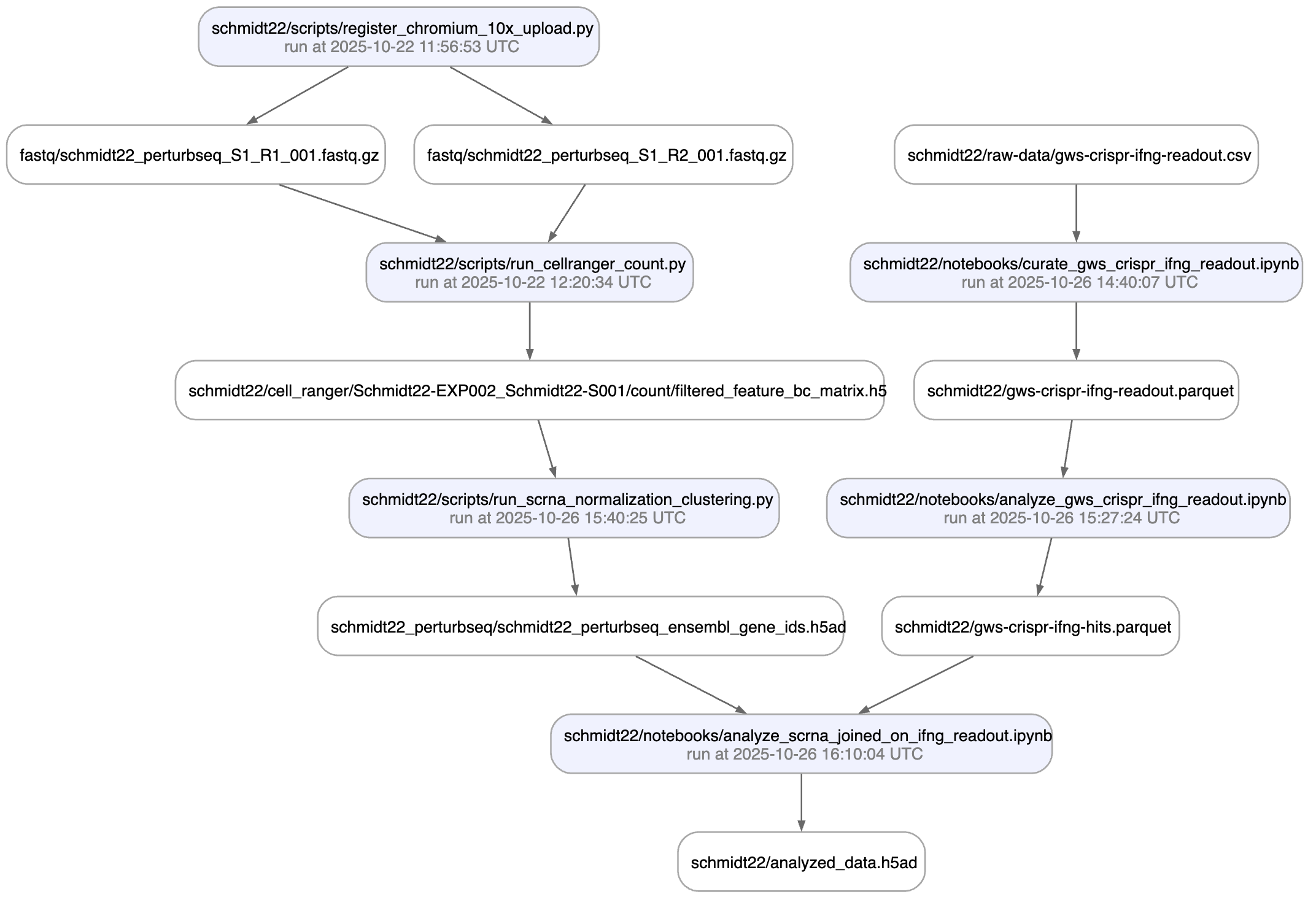
A phenotypic CRISPRa screening result is integrated with scRNA-seq data. Here is the result of the screen input:
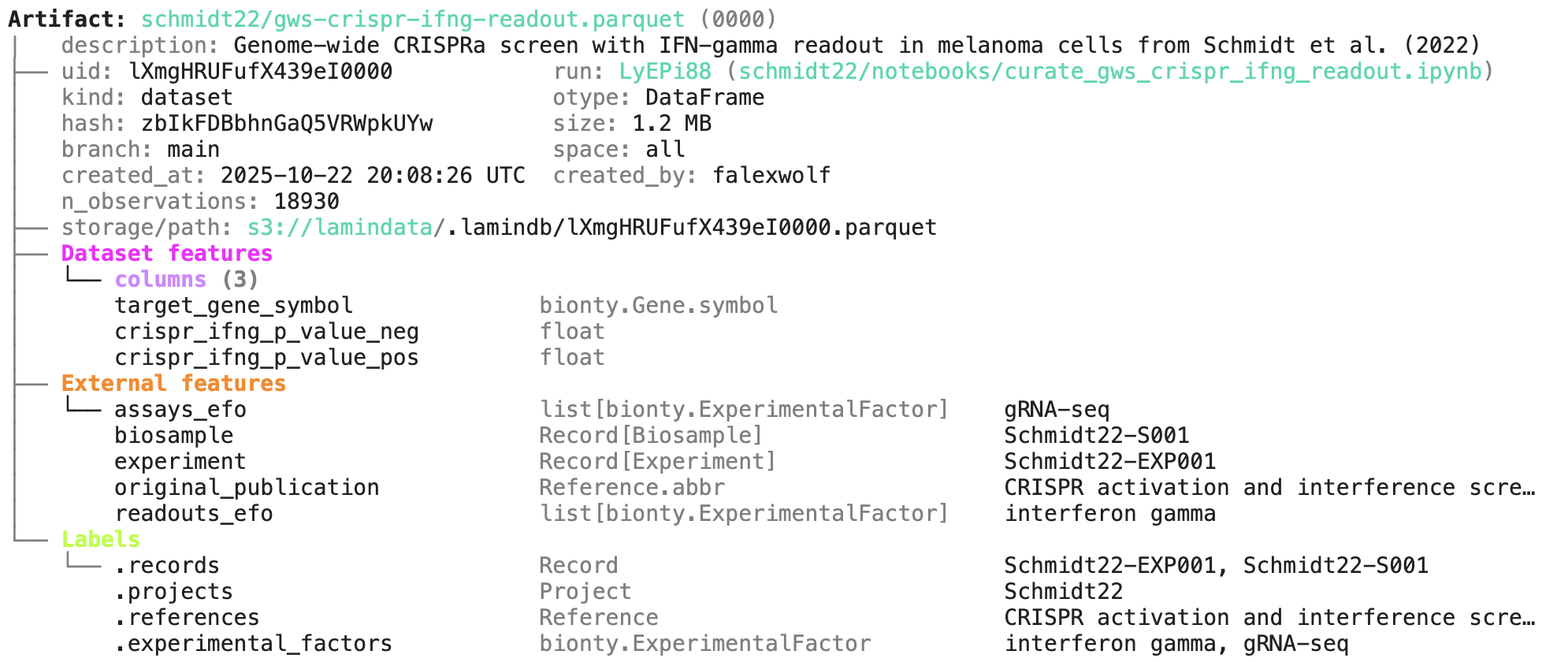
You can label an artifact by running:
my_label = ln.ULabel(name="My label").save() # a universal label
project = ln.Project(name="My project").save() # a project label
artifact.ulabels.add(my_label)
artifact.projects.add(project)Query for it:
ln.Artifact.filter(ulabels=my_label, projects=project).to_dataframe()You can also query by the metadata that lamindb automatically collects:
ln.Artifact.filter(run=run).to_dataframe() # by creating run
ln.Artifact.filter(transform=transform).to_dataframe() # by creating transform
ln.Artifact.filter(size__gt=1e6).to_dataframe() # size greater than 1MBIf you want to include more information into the resulting dataframe, pass include.
ln.Artifact.to_dataframe(include=["created_by__name", "storage__root"]) # include fields from related registriesNote: The query syntax for DB objects and for your default database is the same.
You can annotate datasets and samples with features. Let's define some:
from datetime import date
ln.Feature(name="gc_content", dtype=float).save()
ln.Feature(name="experiment_note", dtype=str).save()
ln.Feature(name="experiment_date", dtype=date, coerce=True).save() # accept date stringsDuring annotation, feature names and data types are validated against these definitions.
artifact.features.set_values({
"gc_content": 0.55,
"experiment_note": "Looks great",
"experiment_date": "2025-10-24",
})Query for it:
ln.Artifact.filter(experiment_date="2025-10-24").to_dataframe() # query all artifacts annotated with `experiment_date`If you want to include the feature values into the dataframe, pass include.
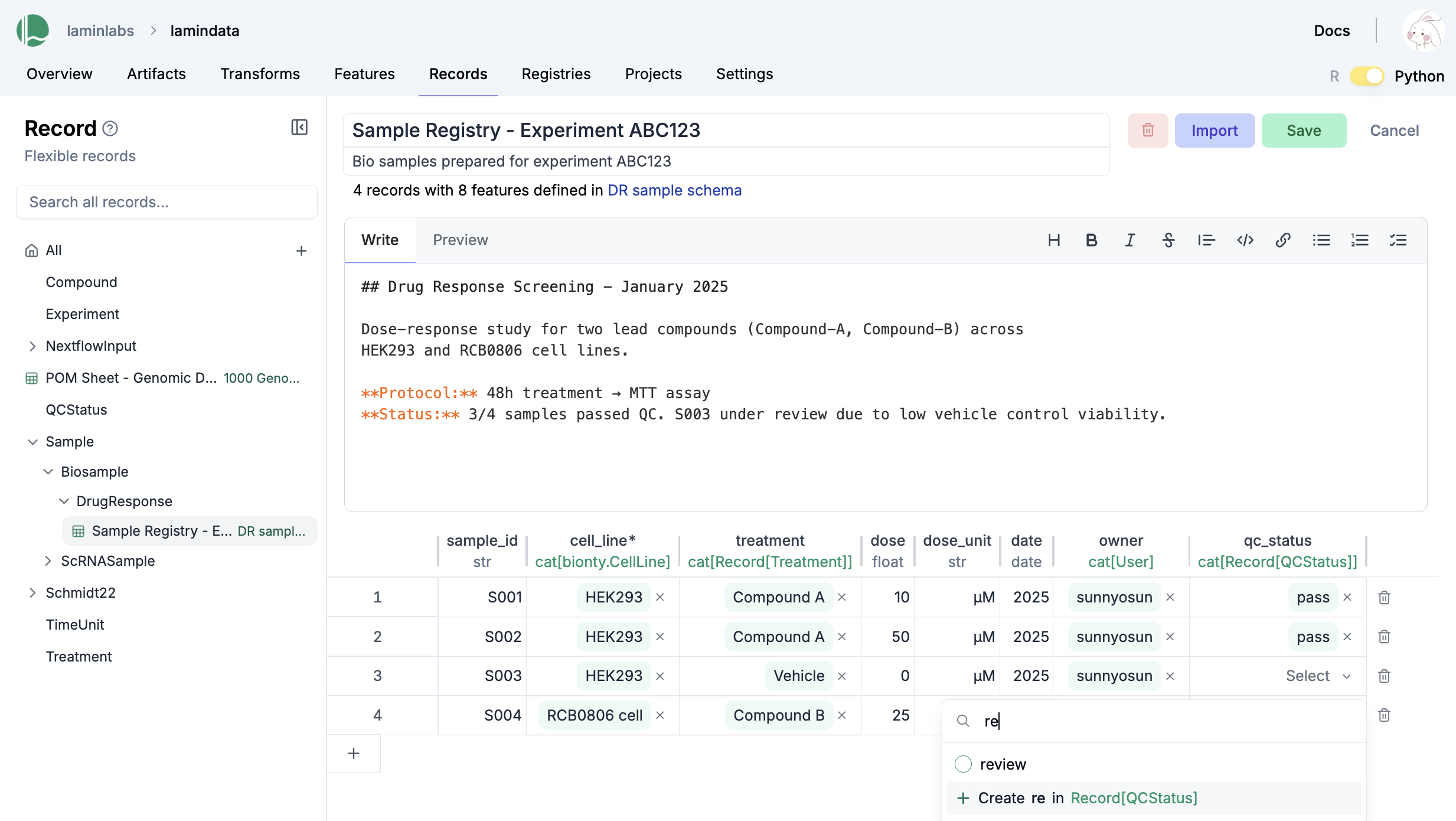
ln.Artifact.to_dataframe(include="features") # include the feature annotationsYou can create records for the entities underlying your experiments: samples, perturbations, instruments, etc., for example:
ln.Record(name="Sample 1", features={"gc_content": 0.5}).save()You can create relationships of entities:
# create a flexible record type to track experiments
experiment_type = ln.Record(name="Experiment", is_type=True).save()
# create a record of type `Experiment` for your first experiment
ln.Record(name="Experiment 1", type=experiment_type).save()
# create a feature to link experiments in records, dataframes, etc.
ln.Feature(name="experiment", dtype=experiment_type).save()
# create a sample record that links the sample to `Experiment 1` via the `experiment` feature
ln.Record(name="Sample 2", features={"gc_content": 0.5, "experiment": "Experiment 1"}).save()You can convert any record type to dataframe/sheet:
experiment_type.to_dataframe()If you change source code or datasets, LaminDB manages versioning for you.
Assume you run a new version of our create-fasta.py script to create a new version of sample.fasta.
import lamindb as ln
ln.track()
open("sample.fasta", "w").write(">seq1\nTGCA\n") # a new sequence
ln.Artifact("sample.fasta", key="sample.fasta", features={"experiment": "Experiment 1"}).save() # annotate with the new experiment
ln.finish()If you now query by key, you'll get the latest version of this artifact:
artifact = ln.Artifact.get(key="sample.fasta") # get artifact by key
artifact.versions.to_dataframe() # see all versions of that artifactTo create a contribution branch and switch to it, run:
lamin switch -c my_branchTo merge a contribution branch into main, run:
lamin switch main # switch to the main branch
lamin merge my_branch # merge contribution branch into mainRead more: docs.lamin.ai/lamindb.branch.
To share data in a lineage-aware way, sync objects from a source database to your default database:
db = ln.DB("laminlabs/lamindata")
artifact = db.Artifact.get(key="example_datasets/mini_immuno/dataset1.h5ad")
artifact.save()This is zero-copy for the artifact's data in storage. Read more: docs.lamin.ai/sync.
Here is how you ingest a DataFrame:
import pandas as pd
df = pd.DataFrame({
"sequence_str": ["ACGT", "TGCA"],
"gc_content": [0.55, 0.54],
"experiment_note": ["Looks great", "Ok"],
"experiment_date": [date(2025, 10, 24), date(2025, 10, 25)],
})
ln.Artifact.from_dataframe(df, key="my_datasets/sequences.parquet").save() # no validationTo validate & annotate the content of the dataframe, use the built-in schema valid_features:
ln.Feature(name="sequence_str", dtype=str).save() # define a remaining feature
artifact = ln.Artifact.from_dataframe(
df,
key="my_datasets/sequences.parquet",
schema="valid_features" # validate columns against features
).save()
artifact.describe()30 sec video.
You can filter for datasets by schema and then launch distributed queries and batch loading.
To validate an AnnData with built-in schema ensembl_gene_ids_and_valid_features_in_obs, call:
import anndata as ad
import numpy as np
import pandas as pd
adata = ad.AnnData(
X=np.ones((21, 10)),
obs=pd.DataFrame({'cell_type_by_model': ['T cell', 'B cell', 'NK cell'] * 7}),
var=pd.DataFrame(index=[f'ENSG{i:011d}' for i in range(10)])
)
artifact = ln.Artifact.from_anndata(
adata,
key="my_datasets/scrna.h5ad",
schema="ensembl_gene_ids_and_valid_features_in_obs"
).save()
artifact.describe()To validate a SpatialData or any other array-like dataset, you need to construct a Schema. You can do this by composing simple pandera-style schemas: docs.lamin.ai/curate.
Plugin bionty gives you >20 public ontologies as SQLRecord registries. This was used to validate the ENSG ids in the adata just before.
import bionty as bt
bt.CellType.import_source() # import the default ontology
bt.CellType.to_dataframe() # your extensible cell type ontology in a simple registryYou can then create objects, e.g. for labeling, analogous to ULabel, Project, or Record:
t_cell = bt.CellType.get(name="T cell")
artifact.cell_types.add(t_cell)Read more: docs.lamin.ai/manage-ontologies.
30 sec video.
When in your development directory, you can save markdown files as records:
lamin save <topic>/<my-note.md>